Chapter 6 More about Factors
This was part of a impromptu session learning about factors.
# load libraries
library(tidyverse)
library(readxl)
library(janitor)
library(forcats)
smoke_complete <- read_excel("data/smoke_complete.xlsx",
sheet=1,
na="NA")6.1 Making a factor variable out of disease
We’re adding a fourth value, BRCA to our levels here.
6.2 Using the character variable
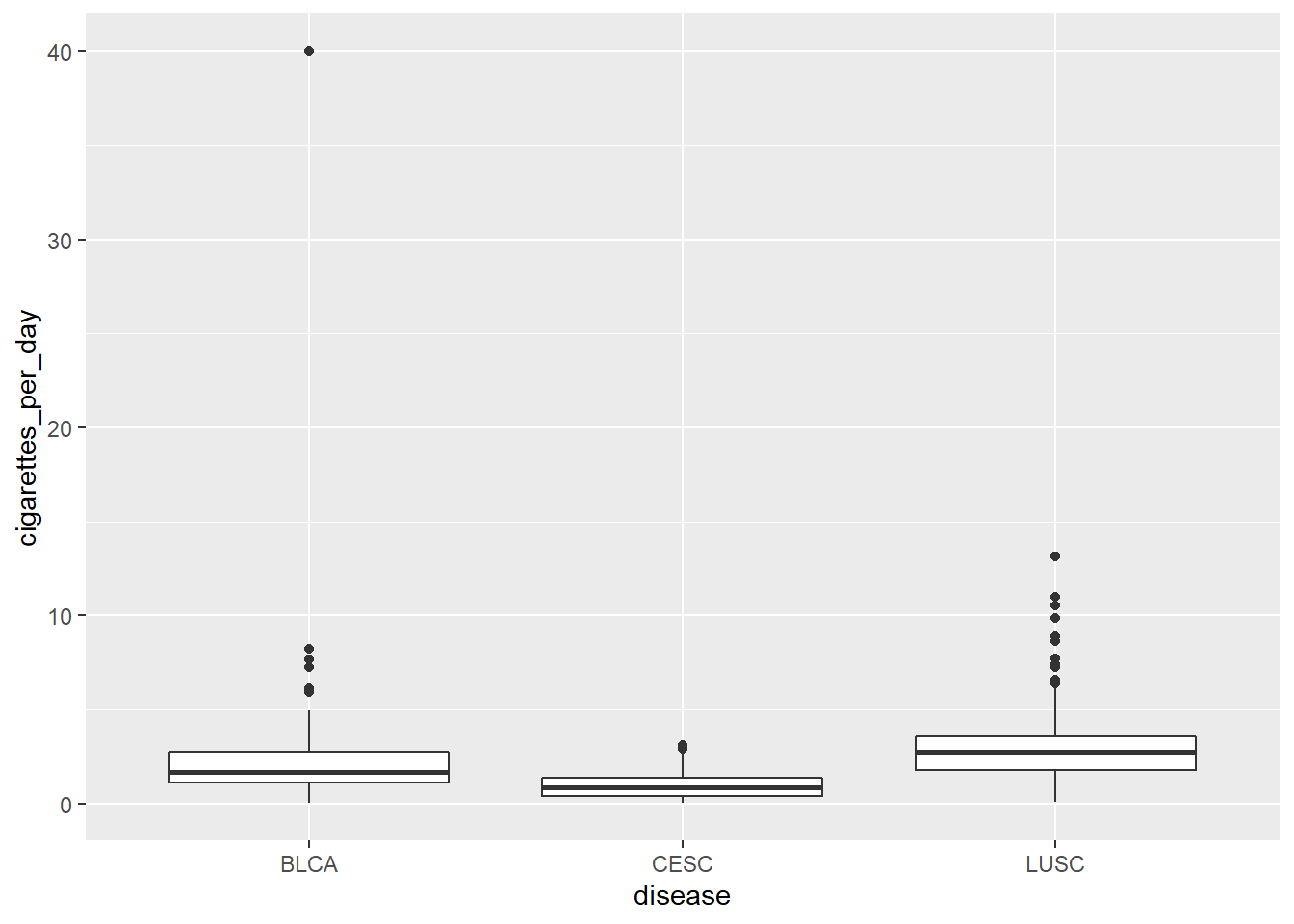
6.3 Compare to the factor variable
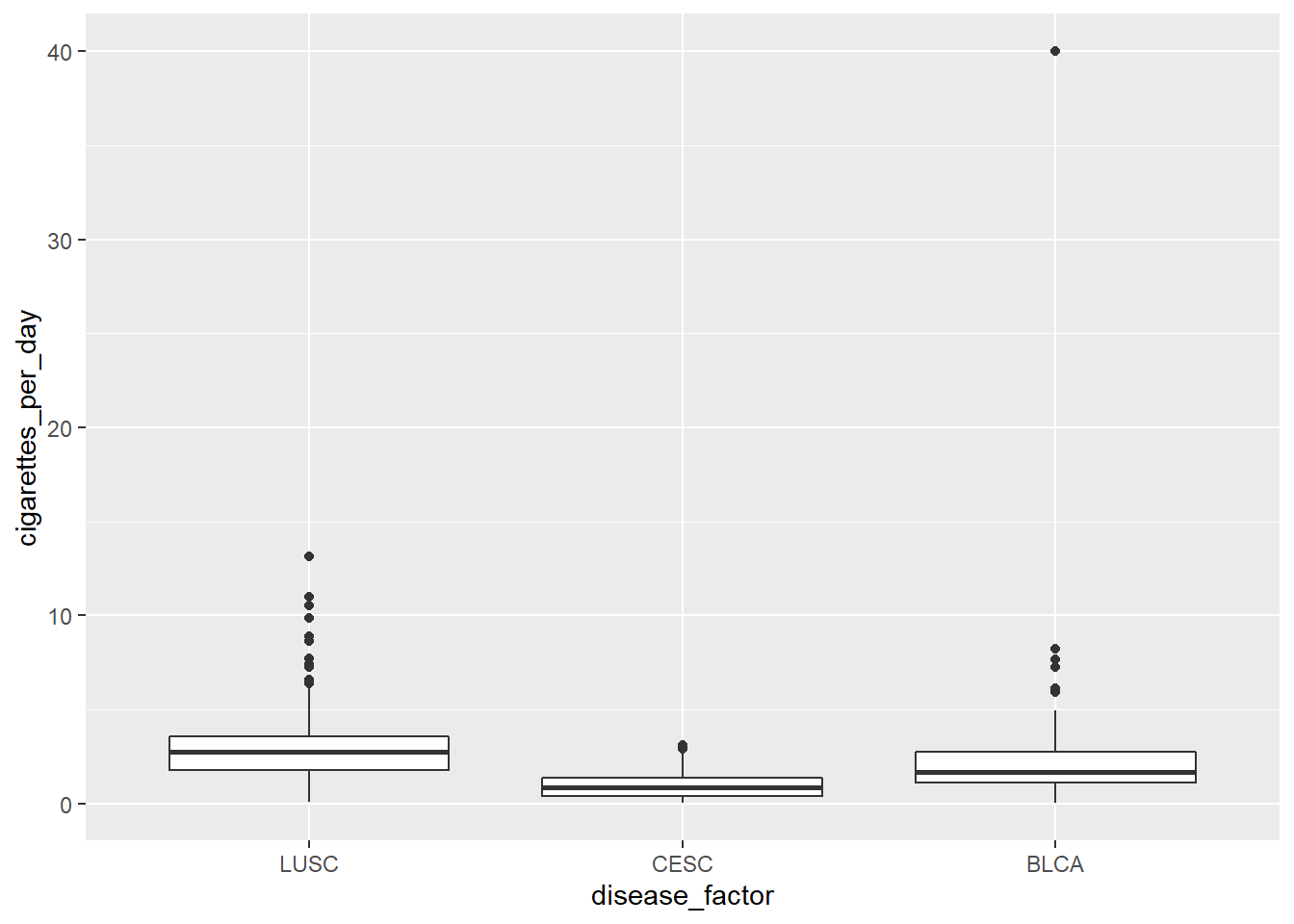
6.4 Another thing about factors
Factor levels also specify the permissible values.
In this example, LUSC and BRCA are the permissible values. We pass a character vector into them, and you can see those values (BLCA, CESC) are recoded as NAs
character_vector <- c("LUSC", "LUSC", "BRCA", "BLCA", "BRCA", "CESC", "CESC")
factor_vector <- factor(character_vector, levels=c("LUSC", "BRCA"))
factor_vector## [1] LUSC LUSC BRCA <NA> BRCA <NA> <NA>
## Levels: LUSC BRCA6.5 fct_rev() - reversing the order of a factor
Very useful when using factors on the y-axis, because the default ordering is first value at the bottom, rather than first value at the top.
library(forcats)
#fct_rev()
smoke_complete3 <- smoke_complete2 %>%
mutate(disease_rev = fct_rev(disease_factor))
#show original factor
ggplot(smoke_complete3) + aes(y=disease_factor, x=cigarettes_per_day) +
geom_boxplot()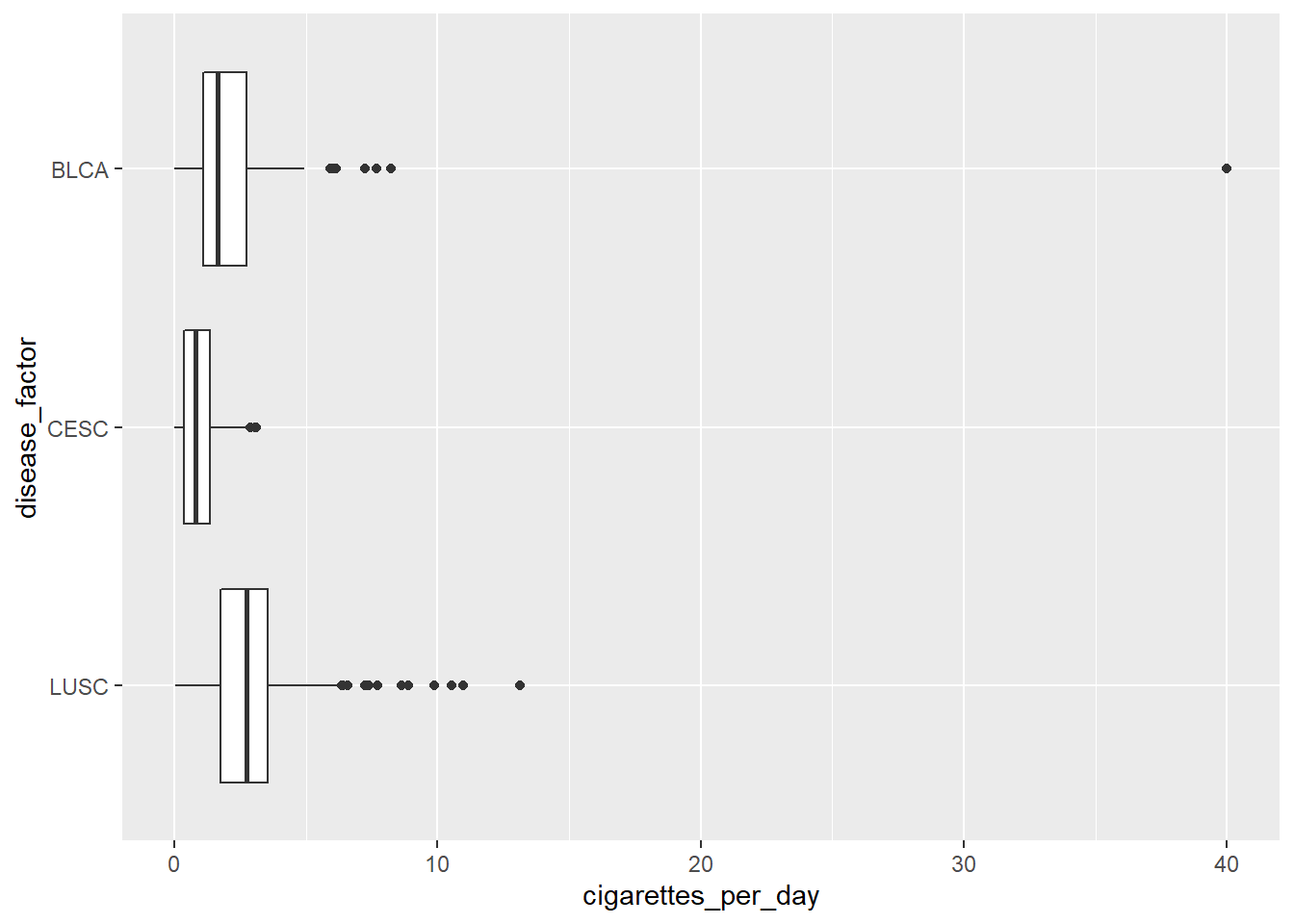
#show factor with reversed order
ggplot(smoke_complete3) + aes(y=disease_rev, x=cigarettes_per_day) +
geom_boxplot()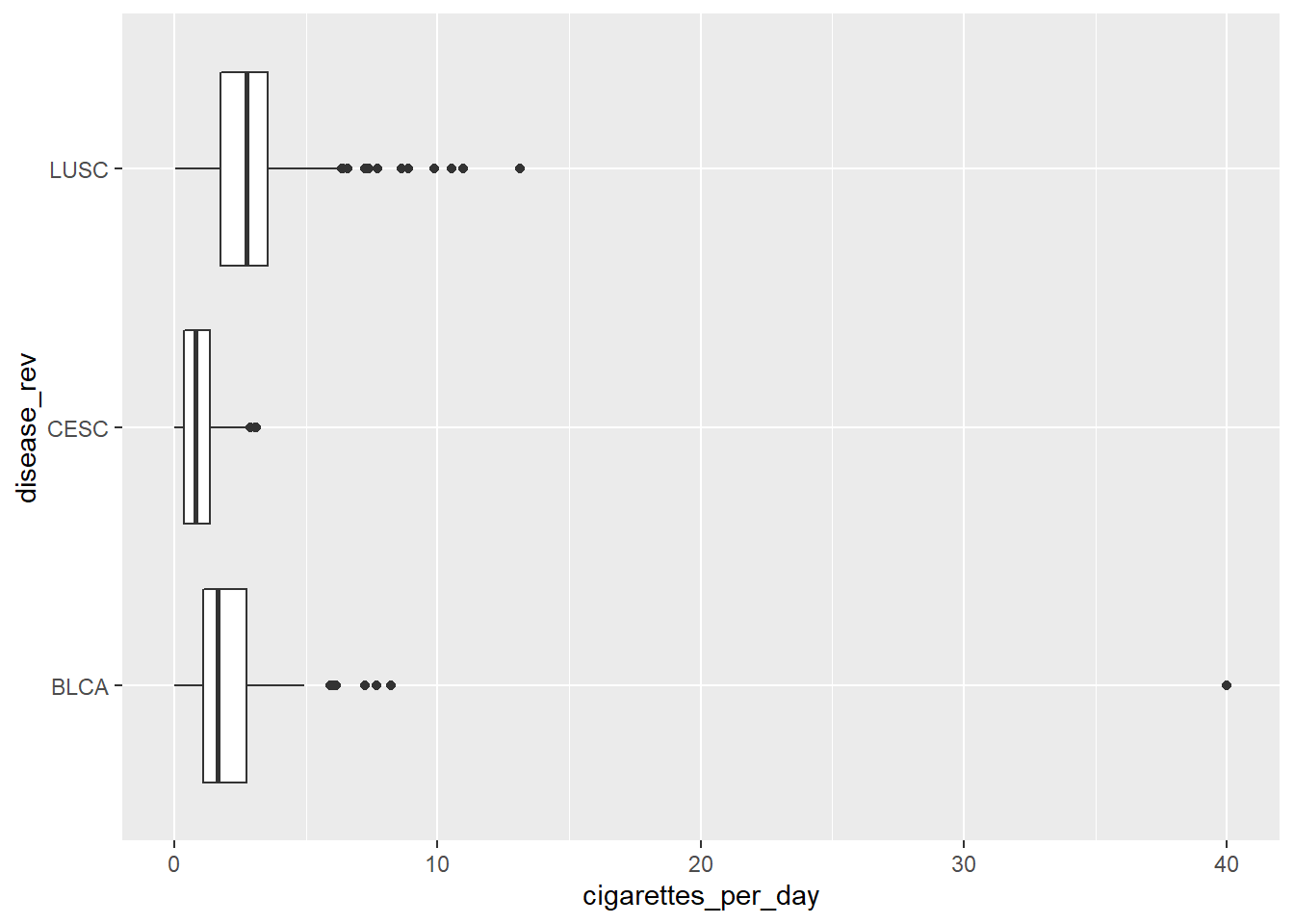
6.6 fct_reorder()
fct_reorder() lets you reorder factors by another numeric variable.
library(forcats)
#fct_rev()
smoke_complete3 <- smoke_complete2 %>%
mutate(disease_reorder = fct_reorder(disease_factor, cigarettes_per_day))
ggplot(smoke_complete3) + aes(y=disease_reorder, x=cigarettes_per_day) +
geom_jitter()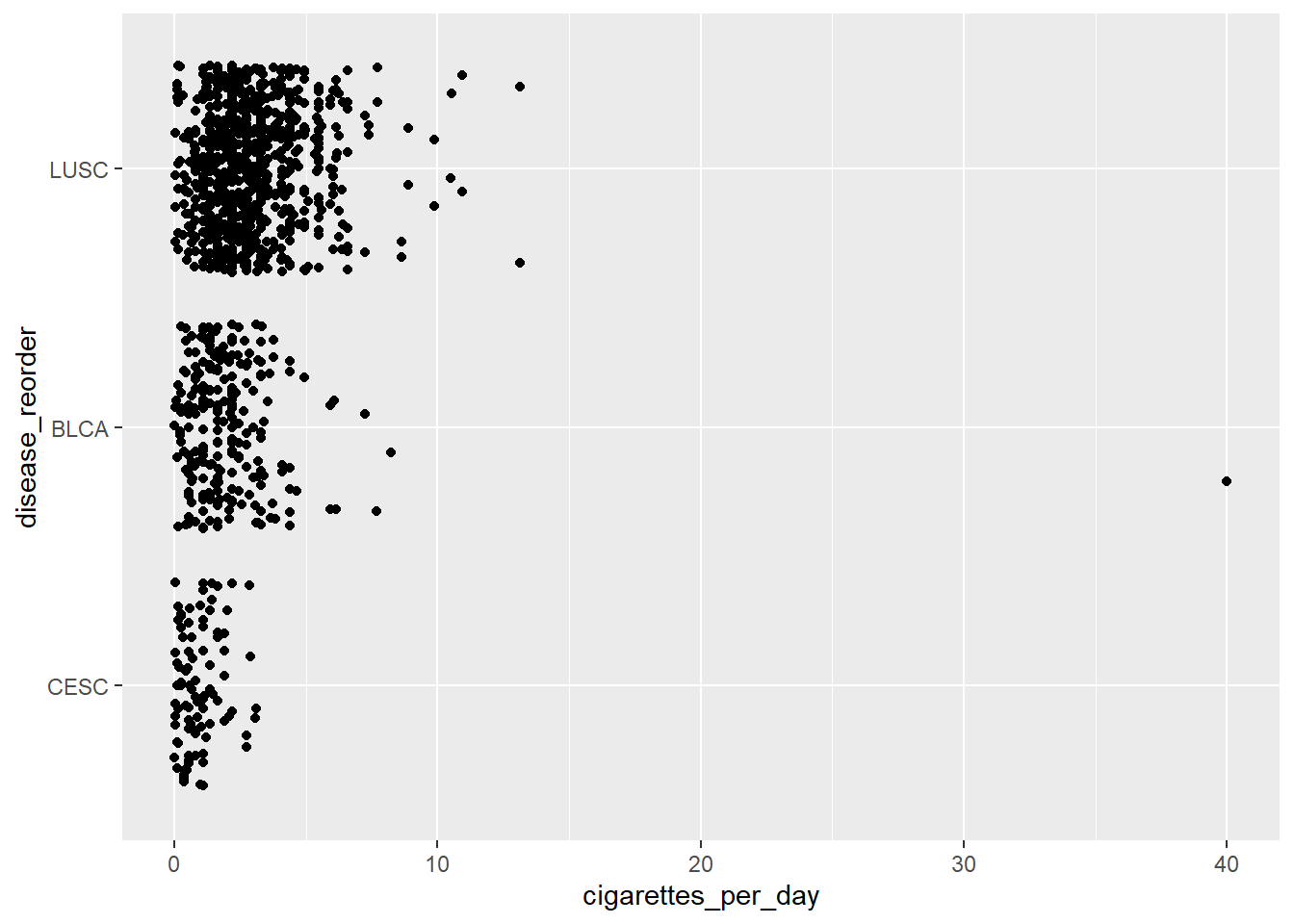
6.7 fct_collapse
fct_collapse() lets you collapse multiple categories into one category.
smoke_complete3 %>%
mutate(disease_collapse = fct_collapse(
disease_factor,
other = c("BLCA", "CESC"),
LUSC = c("LUSC")
)) %>%
tabyl(disease_collapse)## disease_collapse n percent
## LUSC 836 0.7256944
## other 316 0.2743056
## BRCA 0 0.00000006.8 Other really useful forcats functions
fct_recode() - lets you recode values manually.
fct_other() - lets you define what categories are in an other variable.